John Cavanagh, PhD
 Professor & Chair of Biochemistry /Molecular Biology, Associate Dean for Research and Chair of Foundational Sciences
Professor & Chair of Biochemistry /Molecular Biology, Associate Dean for Research and Chair of Foundational Sciences
Department of Biochemistry & Molecular Biology
Office 5E-124B (Laboratories 5S-08 & 5S-18)
Brody School of Medicine
East Carolina University
Greenville, NC 27834
phone: 252-744-2684
email: cavanaghj19@ecu.edu
Education
- University of Surrey, B.S. Chemistry 1985
- University of Cambridge, Ph.D. NMR 1988
- The Scripps Research Institute, Post-Doctoral Fellow 1988-1990
Research Interests
The Cavanagh lab uses a variety of biophysical techniques to study the structures and dynamics of proteins and their targets. By doing this we are able to discern physiological mechanisms of action and points of possible therapeutic intervention. The laboratory’s library of 2-Aminoimidazole compounds has demonstrated efficacy in re-sensitizing multi-drug resistant bacteria to conventional antibiotics. The lab is committed to global health issues, with current investigations focusing on control of protective mechanisms in Bacillus, multi-drug resistance in Acinetobacter baumannii, and regulation of caspase-3 apoptotic activity. The analytical approaches we use include (but aren’t limited to): NMR, X-ray crystallography, computational modeling, mass spectrometry, fluorescence, electron microscopy, surface plasmon resonance.
Selected Publications
Kojetin DJ, Venters RA, Kordys DR, Thompson RJ, Kumar R, Cavanagh J. (2006) Structure, binding interface and hydrophobic transitions of Ca2+-loaded calbindin-D(28K). Nat Struct Mol Biol.
Tucker AT, Bobay BG, Banse AV, Olson AL, Soderblom EJ, Moseley MA, Thompson RJ, Varney KM, Losick R, Cavanagh J. (2014) A DNA mimic: the structure and mechanism of action for the anti-repressor protein AbbA. J Mol Biol.
Feldmann EA, Cavanagh J. (2015) Teaching old drugs new tricks: Addressing resistance in Francisella. Virulence.
Milton ME, Allen CL, Feldmann EA, Bobay BG, Jung DK, Stephens MD, Melander RJ, Theisen KE, Zeng D, Thompson RJ, Melander C, Cavanagh J. (2017) Structure of the Francisella response regulator QseB receiver domain, and characterization of QseB inhibition by antibiofilm 2-aminoimidazole-based compounds. Mol Microbiol.
Milton ME, Minrovic BM, Harris DL, Kang B, Jung D, Lewis CP, Thompson RJ, Melander RJ, Zeng D, Melander C, Cavanagh J. (2018) Re-sensitizing Multidrug Resistant Bacteria to Antibiotics by Targeting Bacterial Response Regulators: Characterization and Comparison of Interactions between 2-Aminoimidazoles and the Response Regulators BfmR from Acinetobacter baumannii and QseB from Francisella spp. Front Mol Biosci.
Palethorpe S, Milton ME, Pesci EC, Cavanagh J. (2022) Structure of the Acinetobacter baumannii PmrA receiver domain and insights into clinical mutants affecting DNA binding and promoting colistin resistance. J Biochem.
Bacterial resistance and Biofilm formation
The World Health Organization cites multi-drug resistant (MDR) bacteria as one of the top three threats to human health. Unfortunately, antibiotic development has essentially ground to a halt thanks to these rapidly emerging resistant strains. No one wants to invest in a therapeutic with such a short shelf life.
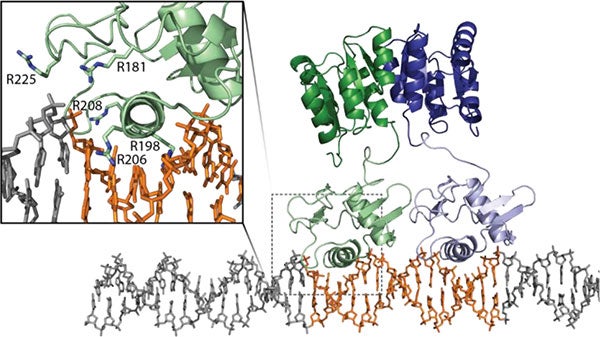 The need for a viable, broad treatment strategy is overwhelming. The only new classes of antibiotics introduced in the last twenty years are active solely against Gram-positive bacteria and resistance to these is already endemic. Additionally, there have been no real advances in treating Gram-negative infections. Without a deeper understanding of the bacterial resistance/persistence and transmission mechanisms, along with the development of new approaches to combat these MDR pathogens, countless people will die.
The need for a viable, broad treatment strategy is overwhelming. The only new classes of antibiotics introduced in the last twenty years are active solely against Gram-positive bacteria and resistance to these is already endemic. Additionally, there have been no real advances in treating Gram-negative infections. Without a deeper understanding of the bacterial resistance/persistence and transmission mechanisms, along with the development of new approaches to combat these MDR pathogens, countless people will die.
We are working to better understand the protective mechanisms that bacteria employ to overcome antibiotics and the host immune response. We have elucidated many functional processes and molecular recognition characteristics of proteins involved in a variety of bacterial responses, including biofilm formation and environmental sensing. These studies were aimed at identifying ‘druggable’ points in signal transduction pathways. As a result of these efforts, in collaboration with the Melander group (University of Notre Dame), we were able to develop an arsenal of compounds (referred to as 2AIs) that re-sensitize MDR bacteria (both Gram-positive and Gram-negative) to current antibiotics.
In addition, the lab is currently working to characterize a complete signaling pathway responsible for biofilm matrix production in order to determine sites for small molecule intervention.
Vancomycin Resistant Enterococci
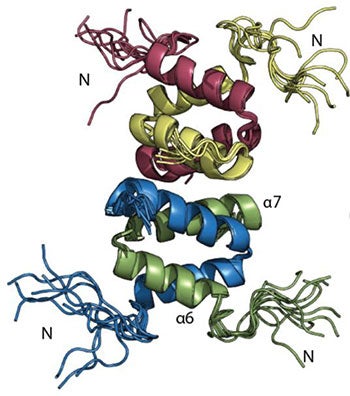 Enterococci are opportunistic pathogens and have emerged as the third leading cause of hospital acquired infection. These infections have become much more insidious by the emergence of MDR strains, making some Enterococci virtually impossible to treat. The worst of the resistant strains are the Vancomycin Resistant Enterococci (VRE) that frequently resist all antibiotics. VRE causes ~33% of Enterococcal infections in ICUs and ~25% of all Enterococci isolated from patients are vancomycin resistant. As part of our ongoing efforts we wish to address the proliferation of VRE. To do this we will focus on E. faecalis V583 – the first VRE strain isolated in the US. To establish infection, VRE uses ethanolamine as a noncompetitive source of carbon/nitrogen, promoting rapid colonization. The ethanolamine sensitive two-component system HK17-RR17 (now EutW-EutV) in E. faecalis V583 has been identified as promoting virulence and survival. Ethanolamine induces a 15-fold increase in the autophosphorylation of the EutW kinase, indicating that this is the signaling ligand recognized by the sensor kinase. This is a remarkable opportunity, since EutW is one of only a dozen histidine kinases whose signaling ligand is known. Additionally, it is also one of the very few fully soluble cytoplasmic kinases and so is more amenable to structure-function studies than its membrane-associated contemporaries. Our primary objective is to establish a pipeline for the structural characterization of the soluble EutW sensor kinase from VRE using NMR, X-ray crystallography, mass spectrometry and molecular modeling. While simultaneously using our current, established, small molecule approach with the response regulator EutV, we will also begin to establish the appropriateness of EutW as a therapeutic target.
Enterococci are opportunistic pathogens and have emerged as the third leading cause of hospital acquired infection. These infections have become much more insidious by the emergence of MDR strains, making some Enterococci virtually impossible to treat. The worst of the resistant strains are the Vancomycin Resistant Enterococci (VRE) that frequently resist all antibiotics. VRE causes ~33% of Enterococcal infections in ICUs and ~25% of all Enterococci isolated from patients are vancomycin resistant. As part of our ongoing efforts we wish to address the proliferation of VRE. To do this we will focus on E. faecalis V583 – the first VRE strain isolated in the US. To establish infection, VRE uses ethanolamine as a noncompetitive source of carbon/nitrogen, promoting rapid colonization. The ethanolamine sensitive two-component system HK17-RR17 (now EutW-EutV) in E. faecalis V583 has been identified as promoting virulence and survival. Ethanolamine induces a 15-fold increase in the autophosphorylation of the EutW kinase, indicating that this is the signaling ligand recognized by the sensor kinase. This is a remarkable opportunity, since EutW is one of only a dozen histidine kinases whose signaling ligand is known. Additionally, it is also one of the very few fully soluble cytoplasmic kinases and so is more amenable to structure-function studies than its membrane-associated contemporaries. Our primary objective is to establish a pipeline for the structural characterization of the soluble EutW sensor kinase from VRE using NMR, X-ray crystallography, mass spectrometry and molecular modeling. While simultaneously using our current, established, small molecule approach with the response regulator EutV, we will also begin to establish the appropriateness of EutW as a therapeutic target.
Small compounds as cancer adjuvants
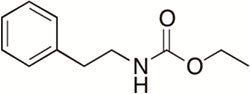 Cytotoxic drugs are the main therapy for combating tumors. Metabolites that have cytotoxic properties have the potential to be lead drugs for cancer treatments. Unfortunately, many cytotoxic compounds are toxic at higher concentrations. This results in a narrow therapeutic index. Lowering the dose of a cytotoxic compound by using it in combination with another active molecule could significantly reduce or eliminate the undesirable toxic side effects. Ethyl N-(2-phenethyl) carbamate is a marine bacterial metabolite that is able to inhibit biofilms in a number of bacterial species It is even more effective at biofilm inhibition when coupled with antibiotics. This observation triggered a study that examined this compound’s ability to enhance other cytotoxic compounds. Two natural products from aquatic microbes, euglenophycin and glycolipid 652, have been shown to have anti-cancer activities and are being assessed for use as chemotherapies. These compounds were found to have significantly magnified cytotoxicity effects in the presence of ethyl N-(2-phenethyl) carbamate analogs. This suggests that lower doses of the anti-cancer drugs could be used to achieve the same effects This would mean that patients receiving chemotherapy treatments could be exposed to a lower dosage, thus reducing the toxic side effects of the treatment. We are currently deciphering the mechanism of action of our compounds.
Cytotoxic drugs are the main therapy for combating tumors. Metabolites that have cytotoxic properties have the potential to be lead drugs for cancer treatments. Unfortunately, many cytotoxic compounds are toxic at higher concentrations. This results in a narrow therapeutic index. Lowering the dose of a cytotoxic compound by using it in combination with another active molecule could significantly reduce or eliminate the undesirable toxic side effects. Ethyl N-(2-phenethyl) carbamate is a marine bacterial metabolite that is able to inhibit biofilms in a number of bacterial species It is even more effective at biofilm inhibition when coupled with antibiotics. This observation triggered a study that examined this compound’s ability to enhance other cytotoxic compounds. Two natural products from aquatic microbes, euglenophycin and glycolipid 652, have been shown to have anti-cancer activities and are being assessed for use as chemotherapies. These compounds were found to have significantly magnified cytotoxicity effects in the presence of ethyl N-(2-phenethyl) carbamate analogs. This suggests that lower doses of the anti-cancer drugs could be used to achieve the same effects This would mean that patients receiving chemotherapy treatments could be exposed to a lower dosage, thus reducing the toxic side effects of the treatment. We are currently deciphering the mechanism of action of our compounds.
Controlling apoptosis
Apoptosis is triggered via two main signaling pathways. The intrinsic pathway initiates from within the cell by developmental cues or as a result of severe cell stress (e.g. DNA damage). The extrinsic pathway initiates when a pro-apoptotic ligand binds to pro-apoptotic receptors on the cell surface. The pathways converge at the level of the effector/executioner caspases (caspases 3, 6 and 7). Once activated, these intracellular enzymes destroy the cellular proteins that are essential for cell survival and trigger cell death.
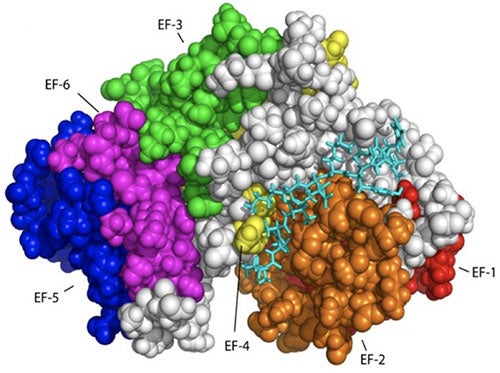 As noted, cancer cells have faulty apoptotic pathways, leading to cell proliferation and the development of malignant tumors. The foremost therapeutic goal in cancer is to re-sensitize tumor cells to apoptosis. Consequently, it is important to define a target that (i) has significant, well-established, anti-apoptotic function in the cell and (ii) is known to be expressed at higher levels in cancerous cells than normal cells. Considering all of these points, the anti-apoptotic calcium-binding protein calbindin-D28K is an especially appealing target. It is well known that small cell lung cancer and non-small cell lung cancer cells have significantly higher concentration of calbindin-D28K compared to normal lung extracts. In addition, the non-small cell lung cancer cells in these studies are also seen to appear at an advanced cancer stage High expression levels of calbindin-D28K have also been observed in medulloblastomas, a highly malignant brain tumor predominantly affecting children under 10 years of age. Overexpression of calbindin-D28K protects osteocytics and osteoblastic (bone) cells from induced apoptosis. Furthermore, a human endometrial adenocarcinoma cell line that overexpresses calbindin-D28K was also shown to inhibit cell death and decrease the amount of Bax, p53 and caspase-3 expression during H2O2-induced apoptosis.
As noted, cancer cells have faulty apoptotic pathways, leading to cell proliferation and the development of malignant tumors. The foremost therapeutic goal in cancer is to re-sensitize tumor cells to apoptosis. Consequently, it is important to define a target that (i) has significant, well-established, anti-apoptotic function in the cell and (ii) is known to be expressed at higher levels in cancerous cells than normal cells. Considering all of these points, the anti-apoptotic calcium-binding protein calbindin-D28K is an especially appealing target. It is well known that small cell lung cancer and non-small cell lung cancer cells have significantly higher concentration of calbindin-D28K compared to normal lung extracts. In addition, the non-small cell lung cancer cells in these studies are also seen to appear at an advanced cancer stage High expression levels of calbindin-D28K have also been observed in medulloblastomas, a highly malignant brain tumor predominantly affecting children under 10 years of age. Overexpression of calbindin-D28K protects osteocytics and osteoblastic (bone) cells from induced apoptosis. Furthermore, a human endometrial adenocarcinoma cell line that overexpresses calbindin-D28K was also shown to inhibit cell death and decrease the amount of Bax, p53 and caspase-3 expression during H2O2-induced apoptosis.
Considering all of these points, calbindin-D28K is an especially appealing target. The general goals of this project are to accurately define how calbindin D28k modulates caspase 3 and to develop small molecules that will remove the ability of calbindin D28K to affect caspase 3, thereby enabling apoptosis to occur.
Cavanagh Lab

Felicia Jaimes, John Cavanagh, Dr. Morgan Milton, Richele Thompson, Sam Palethorpe